Difference between revisions of "Main Page"
Jump to navigation
Jump to search
| Line 7: | Line 7: | ||
Model summary: [[MEDIA:summary.txt|summary]] | Model summary: [[MEDIA:summary.txt|summary]] | ||
| − | The automatic reconstruction of '' | + | The automatic reconstruction of ''Chondrus crispus'' results to a Genome scale [[MEDIA:chondrus_1.3.sbml|Model]] containing 2038 reactions, 2207 metabolites, 2012 genes and 1110 pathways. This GeM was obtained based on the following sources: |
* Based on orthology data: | * Based on orthology data: | ||
** Creation of a global metabolic network containing 1732 reactions | ** Creation of a global metabolic network containing 1732 reactions | ||
Revision as of 13:56, 21 October 2019
CCRGEM-1.3 description
CcrGEM is a genome-scale metabolic network for the model red alga Chondrus crispus.
For more information about this project, please contact Gabriel Markov (Station Biologique de Roscoff).
Automatic reconstruction with AuReMe
Download AuReMe Input Model summary: summary
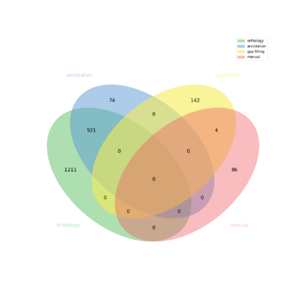
The automatic reconstruction of Chondrus crispus results to a Genome scale Model containing 2038 reactions, 2207 metabolites, 2012 genes and 1110 pathways. This GeM was obtained based on the following sources:
- Based on orthology data:
- Creation of a global metabolic network containing 1732 reactions
- From template ectocarpus_siliculosus creation of a metabolic network containing: 1161 reactions
- From template galdieria.sulphuraria creation of a metabolic network containing: 1361 reactions
- From template arabidopsis_thaliana creation of a metabolic network containing: 383 reactions
- Creation of a global metabolic network containing 1732 reactions
- Based on annotation data:
- Creation of a metabolic network containing 595 reactions
- Based on gap-filling:
- 146 reaction(s) added
- Based on expertise:
- 90 reaction(s) added
- List of tools used:
- Tool: Pantograph
- Tool: PathwayTools
- Tool: Meneco