Difference between revisions of "CPD-10792"
Jump to navigation
Jump to search
(Created page with "Category:metabolite == Metabolite CPD-12116 == * common-name: ** demethylmenaquinol-6 * smiles: ** cc(=cccc(c)=cccc(c)=cccc(c)=cccc(c)=cccc(c)=ccc1(c=c(o)c2(c=cc=cc(c(o)=1...") |
(Created page with "== description == == Automatic reconstruction with [http://aureme.genouest.org AuReMe] == Model summary: summary Download '''AuReMe''' Input/Output [LI...") |
||
| Line 1: | Line 1: | ||
| − | [[ | + | == description == |
| − | + | == Automatic reconstruction with [http://aureme.genouest.org AuReMe] == | |
| − | * | + | Model summary: [[MEDIA:summary.txt|summary]] |
| − | ** | + | |
| − | * | + | Download '''AuReMe''' Input/Output [LINK OR MEDIA data] |
| − | ** | + | |
| − | * | + | The automatic reconstruction of ''MODEL_NAME'' results to a Genome scale [[MEDIA:model.xml|Model]] containing 3356 reactions, 3250 metabolites, 5018 genes and 1391 pathways. This GeM was obtained based on the following sources: |
| − | ** | + | * Based on orthology data: |
| − | * | + | ** Creation of a global metabolic network containing 1879 reactions |
| − | ** | + | *** From template ''ectocarpus_siliculosus'' creation of a metabolic network containing: 1492 reactions |
| − | == | + | *** From template ''arabidopsis_thaliana'' creation of a metabolic network containing: 419 reactions |
| − | * [[ | + | *** From template ''nannochloropsis_salina'' creation of a metabolic network containing: 589 reactions |
| − | + | * Based on annotation data: | |
| − | + | ** Creation of a metabolic network containing 2552 reactions | |
| − | + | * Based on expertise: | |
| − | + | ** 101 reaction(s) added | |
| − | + | * Based on gap-filling: | |
| + | *** 90 reaction(s) added | ||
| + | |||
| + | *List of tools used: | ||
| + | ** Tool: [http://pathtastic.gforge.inria.fr Pantograph] | ||
| + | ** Tool: [http://bioinformatics.ai.sri.com/ptools/ PathwayTools] | ||
| + | |||
| + | ** Tool: [https://pypi.python.org/pypi/meneco Meneco] | ||
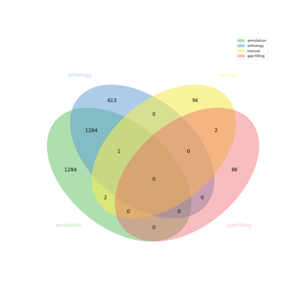
| + | [[FILE:venn.png|frameless|border]] | ||
| + | |||
| + | == Collaborative curation == | ||
| + | * Suggest reactions to add or remove: | ||
| + | ** Download this [[MEDIA:Add_delete_reaction.csv|form]] | ||
| + | * Suggest new reactions to create and add: | ||
| + | ** Download this [[MEDIA:Reaction_creator.csv|form]] | ||
| + | * '''Follow the examples given in the form(s) to correctly share your suggestions''' | ||
| + | * Send the filled form(s) to: CONTACT_MAIL | ||
Revision as of 15:00, 5 January 2021
description
Automatic reconstruction with AuReMe
Model summary: summary
Download AuReMe Input/Output [LINK OR MEDIA data]
The automatic reconstruction of MODEL_NAME results to a Genome scale Model containing 3356 reactions, 3250 metabolites, 5018 genes and 1391 pathways. This GeM was obtained based on the following sources:
- Based on orthology data:
- Creation of a global metabolic network containing 1879 reactions
- From template ectocarpus_siliculosus creation of a metabolic network containing: 1492 reactions
- From template arabidopsis_thaliana creation of a metabolic network containing: 419 reactions
- From template nannochloropsis_salina creation of a metabolic network containing: 589 reactions
- Creation of a global metabolic network containing 1879 reactions
- Based on annotation data:
- Creation of a metabolic network containing 2552 reactions
- Based on expertise:
- 101 reaction(s) added
- Based on gap-filling:
- 90 reaction(s) added
- List of tools used:
- Tool: Pantograph
- Tool: PathwayTools
- Tool: Meneco