Difference between revisions of "Main Page"
Jump to navigation
Jump to search
(Created page with "{{#ask: Category:reaction reconstruction category::orthology | ?common-name | ?ec-number | ?reconstruction tool | ?reconstruction source | ?reconstruction comment | ?n...") |
(Created page with "== description == == Automatic reconstruction with [http://aureme.genouest.org AuReMe] == Model summary: summary Download '''AuReMe''' Input/Output [LI...") |
||
| Line 1: | Line 1: | ||
| − | + | == description == | |
| − | | | + | == Automatic reconstruction with [http://aureme.genouest.org AuReMe] == |
| − | + | Model summary: [[MEDIA:summary.txt|summary]] | |
| − | + | ||
| − | | | + | Download '''AuReMe''' Input/Output [LINK OR MEDIA data] |
| − | + | ||
| − | | | + | The automatic reconstruction of ''Fibrobacter succinogene S85'' results to a Genome scale [[MEDIA:model.xml|Model]] containing 1565 reactions, 1586 metabolites, 1317 genes and 931 pathways. This GeM was obtained based on the following sources: |
| − | + | * Based on annotation data: | |
| − | + | ** Creation of a metabolic network containing 827 reactions | |
| + | * Based on orthology data: | ||
| + | ** Creation of a global metabolic network containing 377 reactions | ||
| + | * Based on expertise: | ||
| + | ** 490 reaction(s) added | ||
| + | * Based on gap-filling: | ||
| + | *** 75 reaction(s) added | ||
| + | |||
| + | *List of tools used: | ||
| + | ** Tool: [http://bioinformatics.ai.sri.com/ptools/ PathwayTools] | ||
| + | ** Tool: [https://github.com/davidemms/OrthoFinder Orthofinder] | ||
| + | |||
| + | ** Tool: [https://pypi.python.org/pypi/meneco Meneco] | ||
| + | [[FILE:venn.png|frameless|border]] | ||
| + | |||
| + | == Collaborative curation == | ||
| + | * Suggest reactions to add or remove: | ||
| + | ** Download this [[MEDIA:Add_delete_reaction.csv|form]] | ||
| + | * Suggest new reactions to create and add: | ||
| + | ** Download this [[MEDIA:Reaction_creator.csv|form]] | ||
| + | * '''Follow the examples given in the form(s) to correctly share your suggestions''' | ||
| + | * Send the filled form(s) to: gem-aureme[at]inria.fr | ||
Latest revision as of 11:09, 17 October 2022
description
Automatic reconstruction with AuReMe
Model summary: summary
Download AuReMe Input/Output [LINK OR MEDIA data]
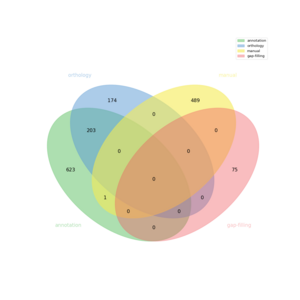
The automatic reconstruction of Fibrobacter succinogene S85 results to a Genome scale Model containing 1565 reactions, 1586 metabolites, 1317 genes and 931 pathways. This GeM was obtained based on the following sources:
- Based on annotation data:
- Creation of a metabolic network containing 827 reactions
- Based on orthology data:
- Creation of a global metabolic network containing 377 reactions
- Based on expertise:
- 490 reaction(s) added
- Based on gap-filling:
- 75 reaction(s) added
- List of tools used:
- Tool: PathwayTools
- Tool: Orthofinder
- Tool: Meneco